Introduction
The choroid plexus (CP) is a highly vascularised structure comprised of a complex network of capillaries that produce cerebrospinal fluid (CSF) via ependymal cells lining the brains ventricles1. The CP serves as a critical interface between the brain and periphery, regulating CSF production, maintaining the blood-CSF barrier, and mediating neuroimmune signalling. Structural and functional alterations of the CP are implicated in early neurodegeneration, particularly in AD, where disrupted CSF dynamics, impaired amyloid clearance, and neuroinflammation contribute to disease progression. Polygenic risk scores for AD (AD-PRS) help quantify cumulative genetic risk across thousands of common variants as a preclinical maker of future AD susceptibility, however, the relationship between AD-PRS and CP structure remains unexplored. Recent work also highlights the importance of cellular context in genetic risk. I.e. different brain cell types contribute distinctively to AD-related pathways, including vascular, glial, and barrier-related processes.2
Aims/Objectives
Here, we adopt a cell-specific PRS framework, integrating genome-wide summary statistics with transcriptional profiles, enabling investigation of genetic risk partitioned by a range of neural cell species. We examined associations between whole-genome AD-PRS, cell-type-specific AD-PRS, and CP volume. We hypothesise that CP volume would be associated with AD-PRS, particularly the ependymal cell-specific PRS. We suggest this will reflect the specific AD-linked gene expression profile of ependymal cells that uniquely feature in the CP epithelial lining.2
Methods
In a UK Biobank (UKBB) sample (n=31,943) we used the CP volume image derived phenotype, derived using T1-weighted MRI and the FreeSurfer pipeline3, which segments the CP within the lateral ventricles. We calculated two PRS:
1. Whole-genome AD-PRS, excluding APOE and major histone compatibility (MHC) regions to focus on genome-wide contributions
beyond the major risk loci and complex linkage disequilibrium structure, respectively.
2. Cell-type-specific AD-PRS, computed for 16 brain and vascular-associated cell types (including ependymal, astrocyte, oligodendrocyte, endothelial, pericyte, and smooth muscle cell classes).1
All AD-PRS were generated using gtx, integrating AD summary statistics with cell-type-enriched transcriptomic profiles drawn from 1 who characterized transcriptional signatures across vascular, immune, and neural cell populations in postmortem human brain tissue. The analytic framework follows our recent study that tests associations between imaging phenotypes and polygenic scores derived both genome-wide and at the cell-type level.4
Results
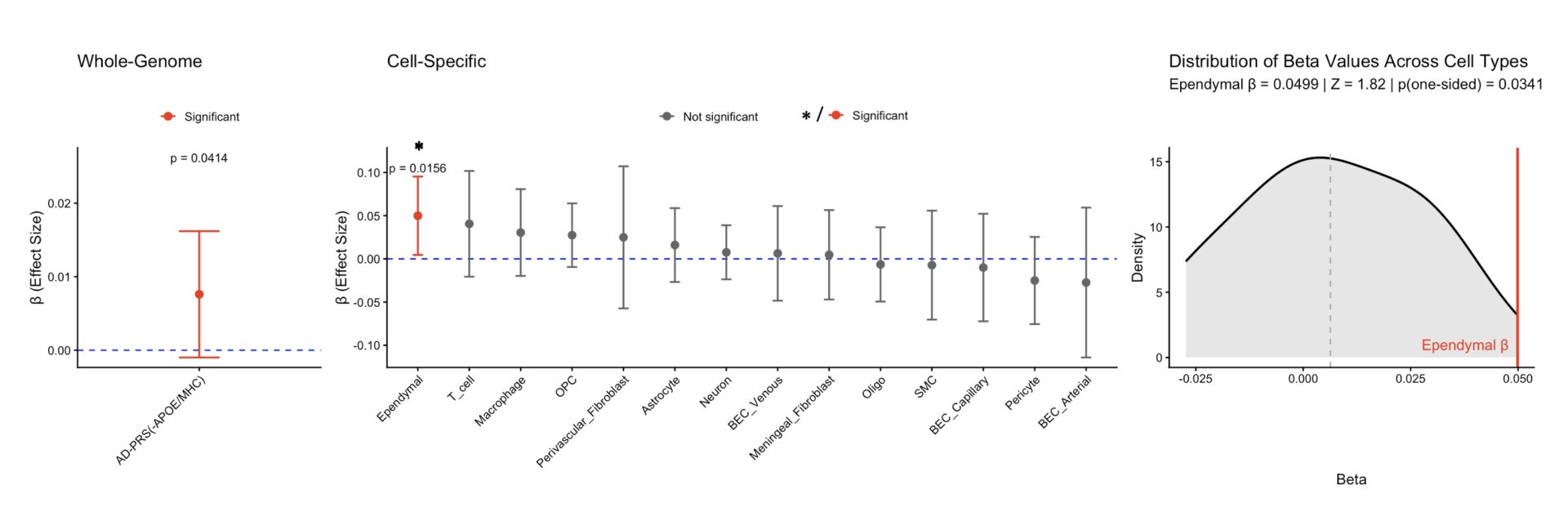
The whole-genome AD-PRS (excluding APOE/MHC regions) was positively associated with CP volume (β=0.0076,SE=0.004,p=0.041, Figure1,left), indicating that global AD-PRS contributes to variation in CP structure. Among cell-specific PRS, ependymal cell AD-PRS showed a robust, highly specific positive association with CP volume (β=0.0499,SE=0.0231,p=0.016, Figure1,center). This association was empirically larger than all other cell-type-specific PRS (Z=1.62,p=0.034, Figure1,right).
Conclusion
Both whole-genome and ependymal cell-specific polygenic risk for AD are associated with CP volume, with the strongest and most specific effect observed for the ependymal PRS. Ependymal cell-specific polygenic risk is selectively and positively associated with CP volume. This highlights a mechanistic link between genetic vulnerability and CP volume, demonstrating cell-type-informed genetic analysis can reveal tissue-specific alterations of AD-related polygenic risk. CP volume represents a promising imaging biomarker for investigating the biological pathways through which genetic risk for AD manifests in the aging brain.
Ethics: Full ethical approval for all data collection by UKBB team.