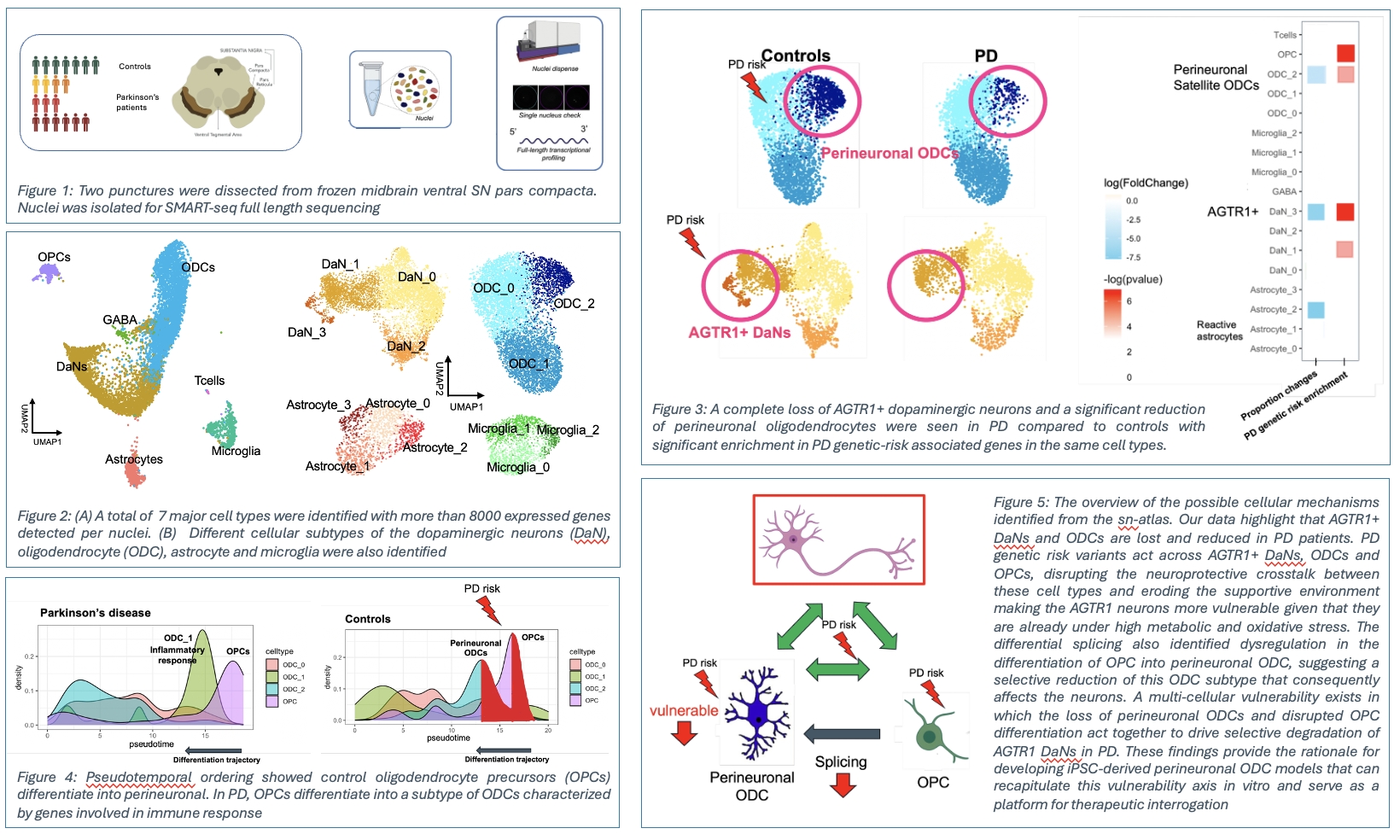
Parkinson’s disease (PD) is a neurodegenerative disorder with incompletely understood cellular and genetic mechanisms. We have created a deep cellular atlas of the human ventral substantia nigra across PD stages by integrating full-length single-nuclei sequencing with genome-wide association studies (GWAS). This atlas revealed that genetic risk converges on AGTR1⁺ dopaminergic neurons and perineuronal oligodendrocytes (pODCs), both of which are reduced in PD, as well as oligodendrocyte precursor cells (OPCs), with risk enriched among disease-disrupted cellular interactions. The atlas also highlighted convergence of PD genetic risk loci with insulin resistance pathways, particularly within pODCs, which showed reduced expression of PI3K/AKT/mTOR pathway genes. Perineuronal oligodendrocytes are a distinct oligodendrocyte population in grey matter that establish direct somatic contact with neuronal cell bodies. Unlike classical oligodendrocytes, these cells do not myelinate, suggesting they function to provide trophic and metabolic support to neurons and modulate neuronal activity. Perineuronal oligodendrocytes are highly understudied.
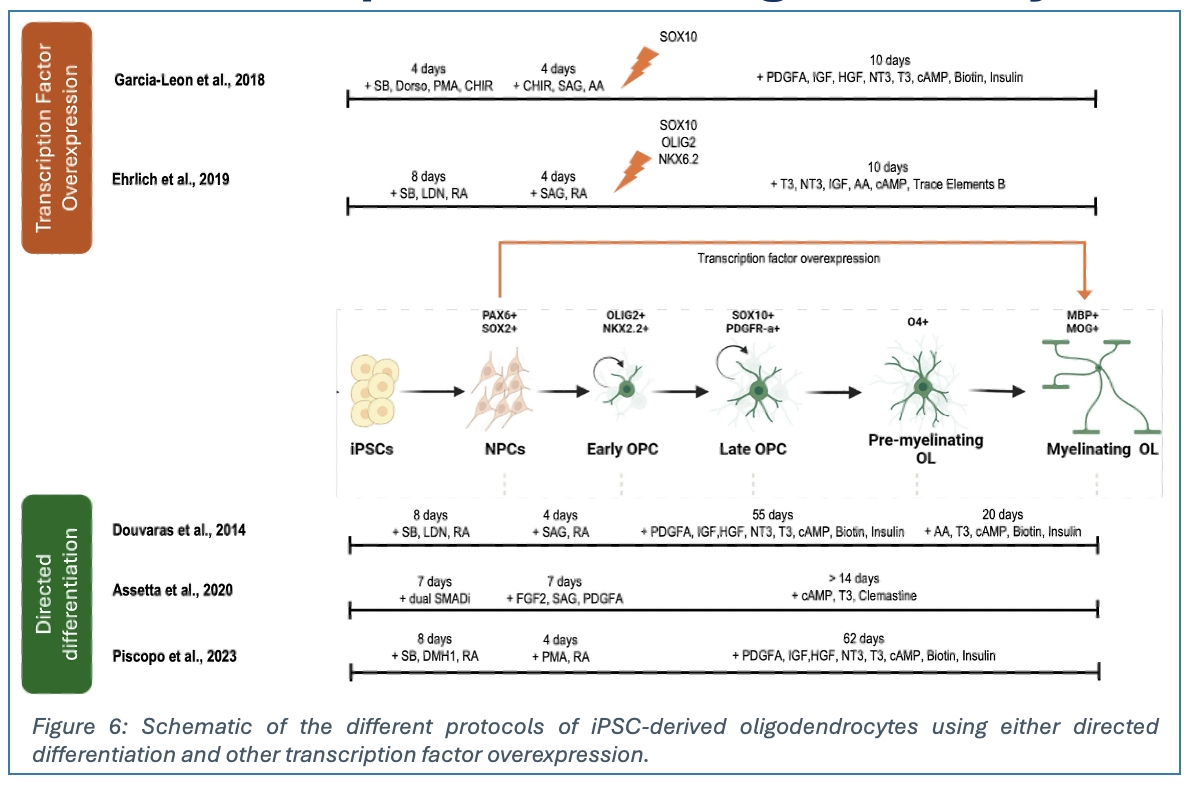
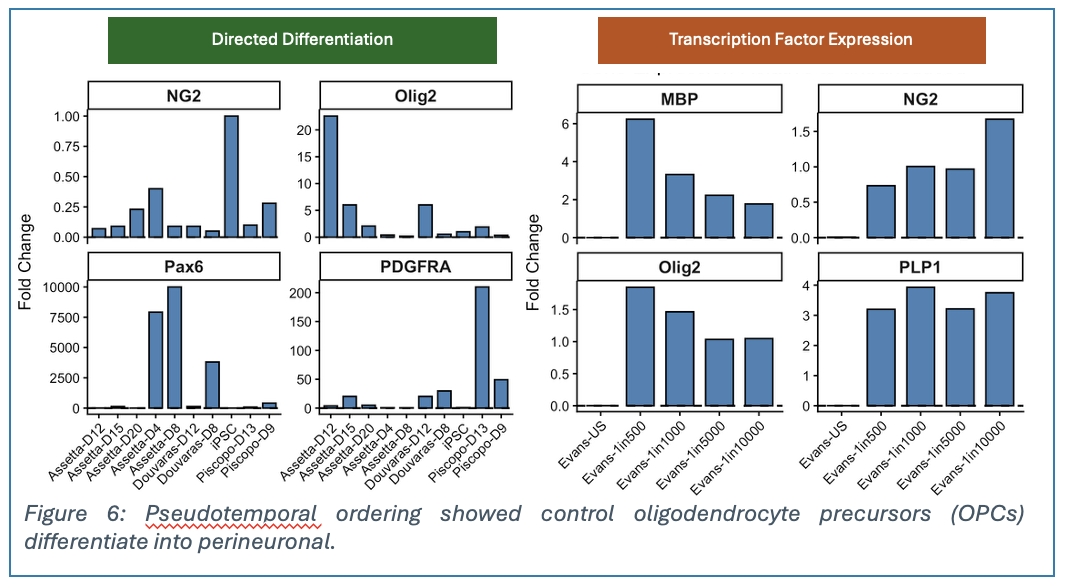
Given our limited understanding of perineuronal oligodendrocytes, we are using cellular atlas insights to guide pODC derivation from induced pluripotent stem cells (iPSCs). We are testing established directed-differentiation protocols to derive OPCs from iPSCs using either neurosphere-based or adherent culture approaches, then differentiate these OPCs into pODCs. We are also testing direct reprogramming methods through overexpression of transcription factor combinations essential for oligodendrocyte fate specification. After establishing robust OPC and oligodendrocyte differentiation, we will introduce modifications to small molecule concentrations, timing, and transcription factor expression patterns obtained from our atlas data and other in silico analysis to promote perineuronal oligodendrocyte fate. A key approach will be to characterise the cells at key differentiation timepoints using scRNA sequencing to track lineage specification. This will be supported by other characterisation methods such as qPCR and immunofluorescence.
Once we establish a robust protocol to derive pODCs, we will increase culture complexity by incorporating pODCs into 3D myelinating organoids (myelinoids) and co-culturing pODCs with iPSC-derived dopaminergic neurons to recapitulate the in vivo perineuronal architecture. This culture system will enable us to investigate PD genetic risk using patient-derived cells and elucidate the metabolic and functional consequences of pODC dysfunction in disease pathogenesis.