Cryptococcus neoformans is a neurotropic environmental fungus causing lethal meningitis in immunocompromised individuals, particularly acquired immunodeficiency syndrome (AIDS) patients. C. neoformans can use microglia as a reservoir for cerebral infection by subverting microglia’s normal immune functions, although the exact phenotypic changes are still poorly described [1]. A growing body of evidence suggests chronic cerebral infections potentiate Alzheimer’s disease (AD), for example, by the fungus Candida albicans. C. albicans infection increases production of amyloid-β, an AD hallmark, as a potential anti-microbial response [2] and drives microglia-mediated neuroinflammation [3]. Mirroring cerebral infections, genome-wide association studies (GWAS) also identified Aβ- and microglia-associated genes as strongly enriched risk factors for late-onset AD [4]. Given C. neoformans is a neurotropic yeast with chronic exposure, and can modulate microglial function, this work explores if C. neoformans could also drive AD-associated mechanisms.
We modelled microglia-C. neoformans interactions in vitro using the human HMC3 microglia cell line infected for 24 hours with the H99E C. neoformans or yeast-locked SC5314-EfgΔ/Δ/Cph1Δ/Δ C. albicans. Fungi were opsonized for 30 minutes in 20% human serum, then inoculated into HMC3 cultures ± 100 U/mL interferon-γ (IFNγ). Microglial viability was assessed via lactate dehydrogenase (LDH) and resazurin assays, for cell lysis and metabolic activity, respectively, while fungal proliferation was determined via a colony forming unit (CFU) assay. Neuroinflammatory responses were measured by enzyme-linked immunosorbent assays (ELISA) for IFNγ, TNFα, and IL-1β. Enrichment of AD pathways in microglia was assessed via RNA sequencing.
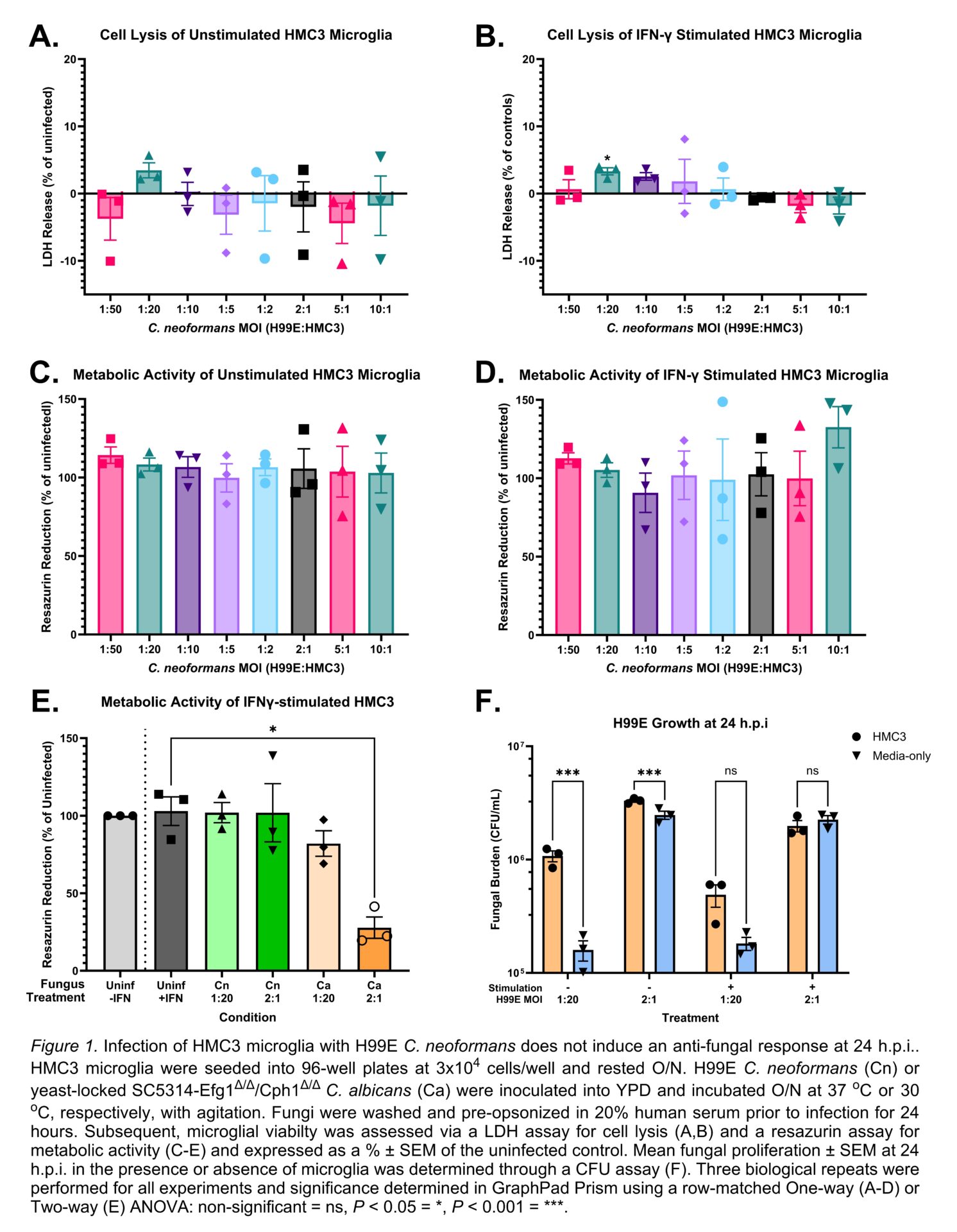
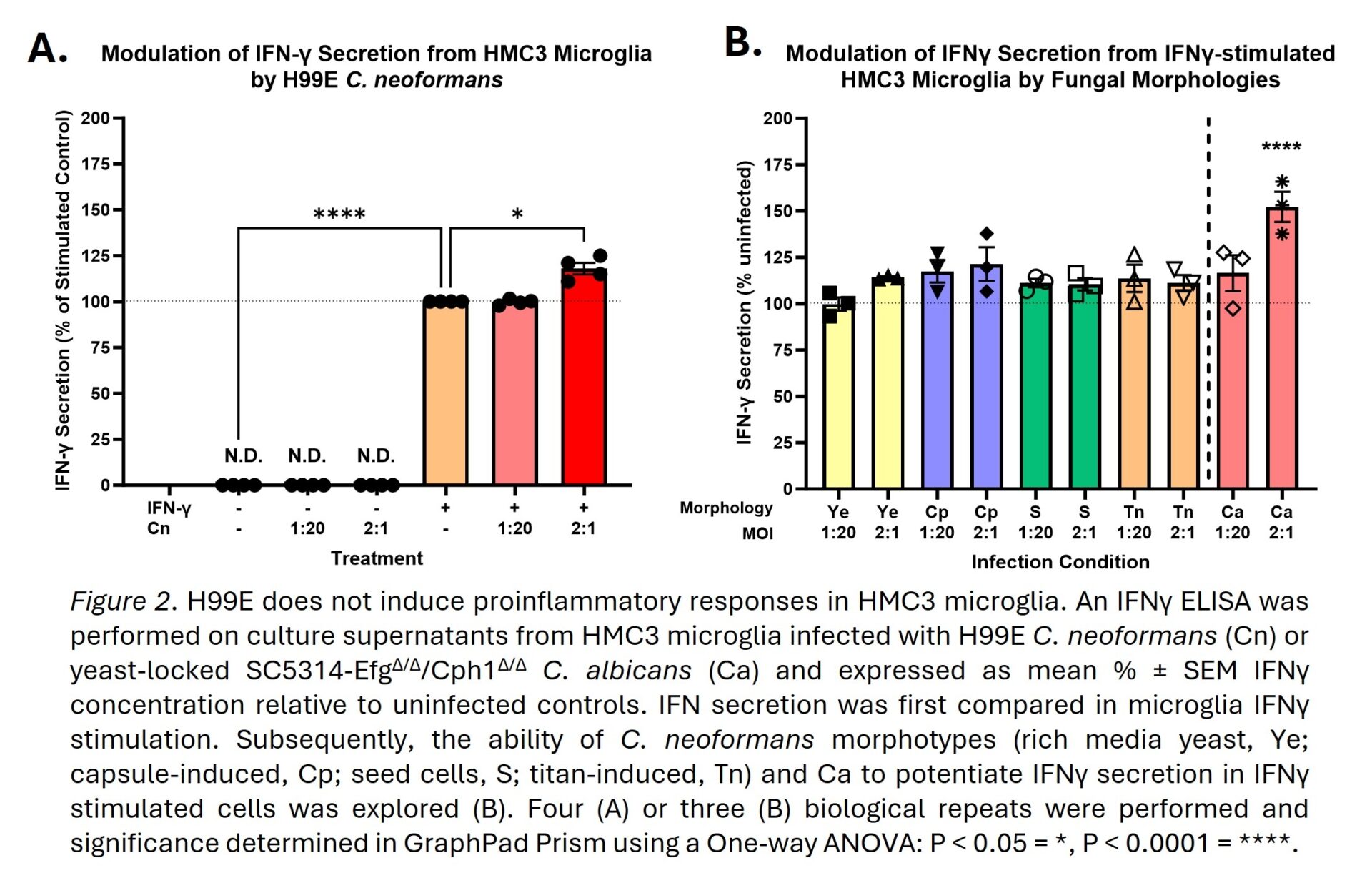
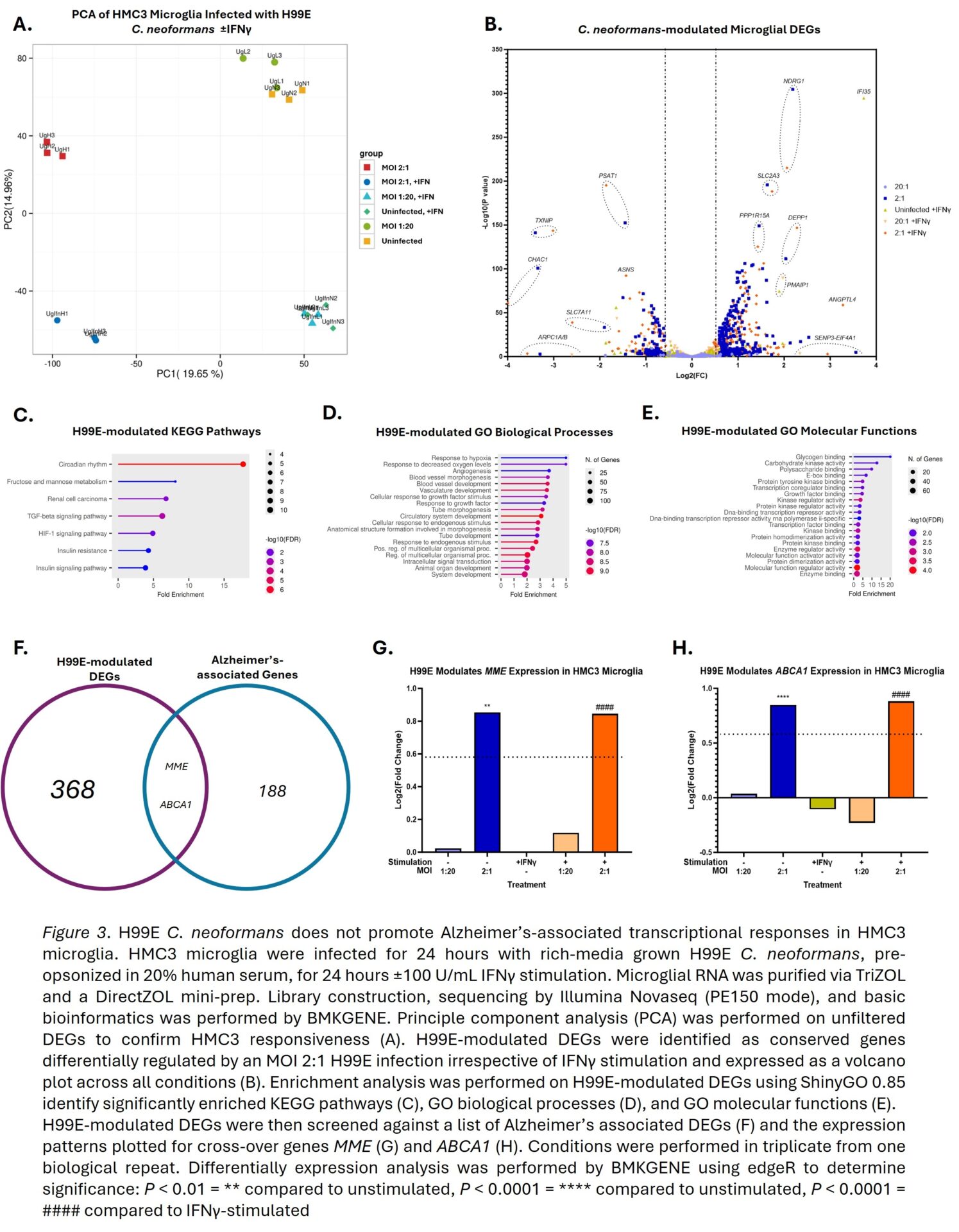
C. neoformans did not increase HMC3 lysis or reduce metabolic activity, figure 1A-D, whilst C. albicans significantly reduced the latter by 72.2%, figure 1E. C. neoformans showed superior replication in the presence of HMC3s compared to media alone, figure 1F, supporting the subversion of microglia antimicrobial responses. C. neoformans only mildly potentiated IFNγ secretion in IFNγ-stimulated HMC3s, figure 2A, and did not induce detectable secretion of TNFα or IL-1β (data not shown). However, when compared to C. albicans, this potentiation became non-significant despite using multiple C. neoformans morphotypes with varying exposure of pathogen-associated molecular patterns (PAMPs), figure 2B. Transcriptomics confirmed HMC3s respond to H99E infection and/or IFNγ treatment, figure 3A, with 368 genes specifically modulated by H99E infection, figure 3B. However, H99E-modulated genes showed no significant enrichment of AD or broader neurodegeneration pathways, figure 3C-E. Additionally, when H99E-modulated genes were crossed-over against 188 AD GWAS, only MME and ABCA1 were conserved, figure 3F, and both genes were regulated in an anti-AD fashion, figure 3G-H.
Overall, our results do not provide any evidence that H99E C. neoformans promotes AD-like mechanisms in a HMC3 microglia model and highlights the strong subversion of antimicrobial responses by C. neoformans. Further investigations into the C. neoformans’ mechanisms of microglial evasion would require more translatable models. For example, we are currently optimizing the use of primary human monocyte-derived microglia-like cells and induced pluripotent stem cell-derived microglia as novel cryptococcal meningitis infection models, figure 4. Additionally, more work should be performed to refine hallmarks of AD-associated microbes for more efficient identification of currently unknown risk factors.
Note: n numbers in figure legends.