Introduction: Alzheimer’s disease (AD) is a neurodegenerative disorder with hallmarks of cerebral β-amyloid (Aβ) plaques, tau hyperphosphorylation, and gliosis (1) leading to cognitive dysfunction (2). Importantly, women account for two-thirds of diagnosed cases (3), present a higher risk of AD (4), and worse clinical manifestation and progression than men (5). Sex-related differences in neuroinflammation and gliosis are suggested to impact AD prevalence and progression. Furthermore, sex differences have been described in the transcriptome, metabolism, morphology, and function of microglia in human AD patients (6) and animal models of the disease (7,8). However, the mechanisms by which these sex-dependant changes impact on AD progression are still unclear.
Aim of study: Previous RNA sequencing studies have investigated sex differences in a case-versus-control manner. We are interested in studying the differences between male and female AD patients to better understand sex-biased alteration of microglial function within the disease.
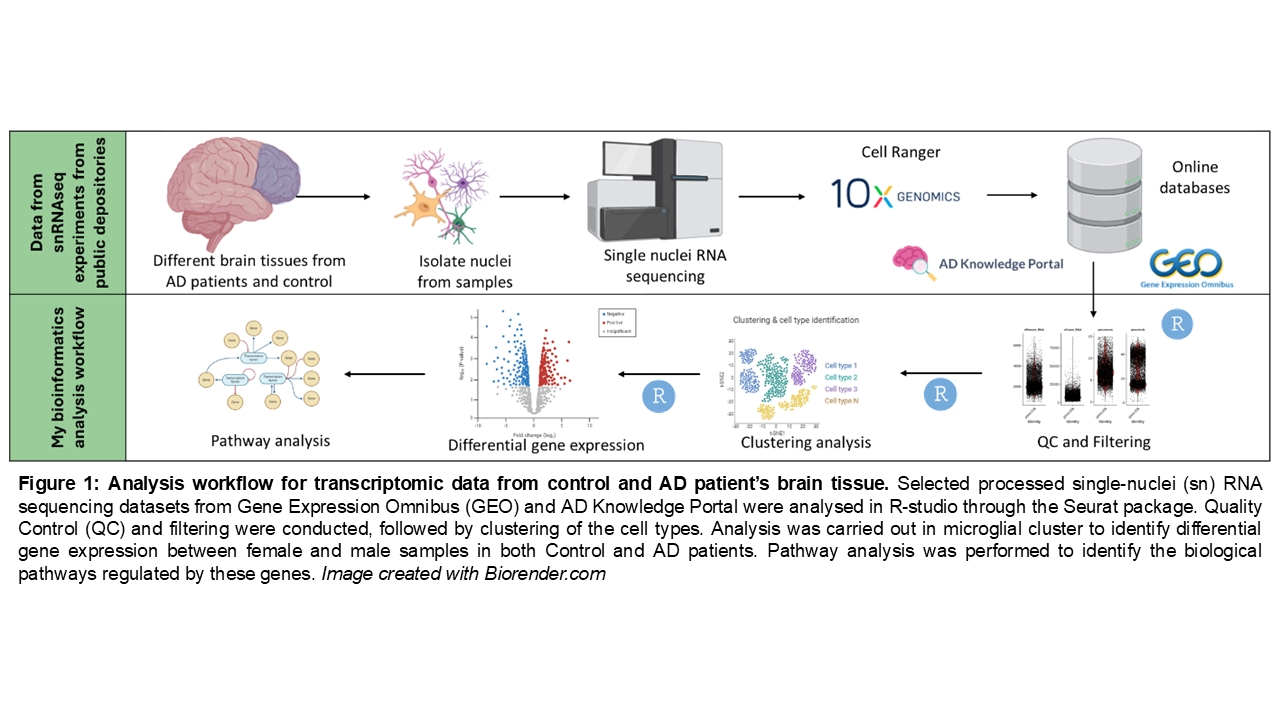
Methods: We utilized publicly available human single-nucleus RNA sequencing datasets containing data from AD patients and non-AD individuals (control). Specifically, we sourced datasets from two repositories: the Gene Expression Omnibus (9) and the AD Knowledge Portal on Synapse (10), selecting those that met our predefined criteria, and carried out our analysis (see Figure 1: workflow of study).
We selected datasets from prefrontal cortex (number of AD samples (M/F) – 9 (6/3), number of control samples (M/F) – 9 (6/3)), dorsolateral prefrontal cortex (number of AD samples (M/F) – 11 (3/8), number of control samples (M/F) – 10 (4/6)), entorhinal cortex (number of AD samples (M/F) – 9 (4/5), number of control samples (M/F) – 9 (5/4)), and somatosensory cortex (number of AD samples (M/F) – 9 (4/5), number of control samples (M/F) – 9 (5/4)). Using RStudio, we performed differential gene expression analysis using the edgeR and MAST statistical methods, comparing microglial cells from male and female patients. To ensure the analysis was AD-specific, we excluded genes that exhibited the same sex-differential expression patterns in control individuals.
Results: We identified genes that present the same sex-biaised expression profile between the two statistical methods, thereby refining understanding of sex-specific differences in microglial gene dysregulation in AD. Specifically, we found that in female AD patients, genes associated with metal binding and homeostasis, cell adhesion, mitochondrial metabolism and structural stabilization were downregulated compared to male AD patients. Interestingly, no genes were commonly dysregulated in female vs male AD patients between all the different brain regions, which suggests the sex-differences are region specific.
Conclusion: Our findings highlight specific microglial functions that may drive sex-specific differences in AD and inform further studies in vitro and in vivo aiming at clarifying biomolecular mechanisms with hopes to guide the development of personalized therapies.